
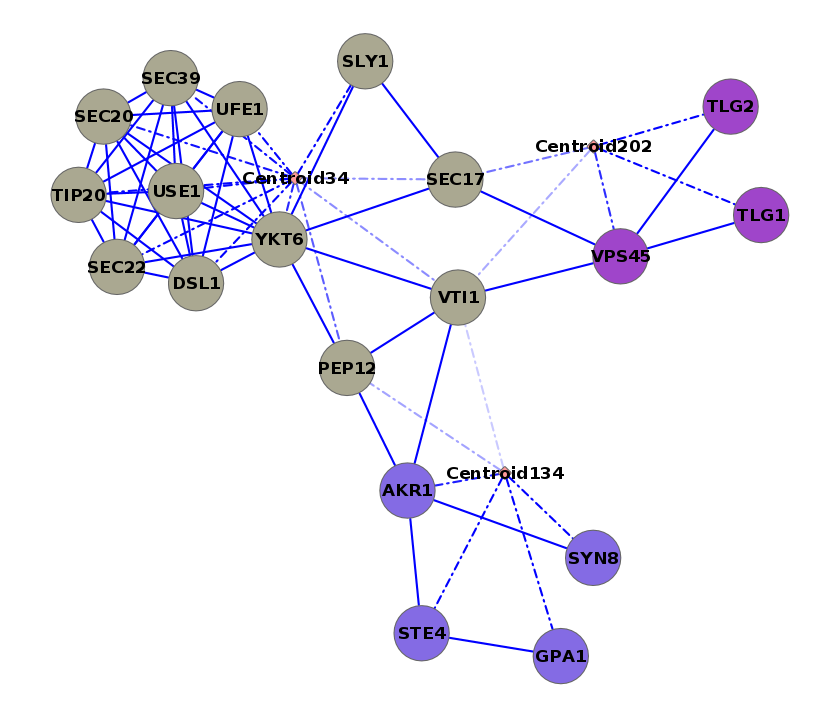
Three complexes are highlighted: SWR1, SET1, and prefoldin. Thick edges represent intra-cluster edges and thin edges are inter-cluster.
(A) The overall results of MCL clustering of the Collins et al., data set showing the largest clusters. In panels A and B, all arrays (cell/tissue types) are horizontally ordered the same as the GSE10246 data set.Ĭlustering of yeast protein-protein interaction networks in the context of overlapping yeast genetic interaction data reveals possible pathway interactions between three well-known complexes. In panel B, all clusters are ordered by decreasing cluster size, and the yellow-cyan color scale is identical to panel A. All expression data are log 2 scaled and median centered. (E) Expression data of four cell/tissue types from GSE10246 superimposed onto the ten largest subnetworks from panel C (Stomach = GSM258771 Lymph Node = GSM258691 Cerebral Cortex = GSM258635 Embryonic Stem Cell = GSM258658). Nodes represent individual cell/tissue types (labeled with first word of each sample name only), node colors correspond to different clusters, and increasing edge thickness and opacity reflect increasing frequency of co-clustering between any given pair of nodes over all ensemble iterations (see ). (D) Fuzzy cluster network of cell/tissue types in GSE10246. (C) Protein interactome divided into subnetworks corresponding to co-expression clusters identified by AutoSOME. Heat maps showing clusters of mouse gene expression data (GSE10246) identified using (A) hierarchical clustering and (B) AutoSOME clustering. Gene expression clustering reveals mouse protein interactome modules and fuzzy relationships among mouse cells and tissues. clusterMaker is available via the Cytoscape plugin manager. Several of these visualizations and algorithms are only available to Cytoscape users through the clusterMaker plugin. The Cytoscape plugin clusterMaker provides a number of clustering algorithms and visualizations that can be used independently or in combination for analysis and visualization of biological data sets, and for confirming or generating hypotheses about biological function. Cytoscape session files for all three scenarios are provided in the Additional Files section. For scenario three, we explore the possible annotation of a protein as a methylmalonyl-CoA epimerase within the VOC superfamily. For scenario two, we explore the prefoldin complex in detail using both physical and genetic interaction clusters. For scenario one, we explore functionally enriched mouse interactomes specific to particular cellular phenotypes and apply fuzzy clustering. Results are presented in the form of three scenarios of use: analysis of protein expression data using a recently published mouse interactome and a mouse microarray data set of nearly one hundred diverse cell/tissue types the identification of protein complexes in the yeast Saccharomyces cerevisiae and the cluster analysis of the vicinal oxygen chelate (VOC) enzyme superfamily.
Cytoscape clustering plus#
clusterMaker is the first Cytoscape plugin to implement such a wide variety of clustering algorithms and visualizations, including the only implementations of hierarchical clustering, dendrogram plus heat map visualization (tree view), k-means, k-medoid, SCPS, AutoSOME, and native (Java) MCL. The Cytoscape network is linked to all of the other views, so that a selection in one is immediately reflected in the others.

Here we present clusterMaker, a Cytoscape plugin that implements several clustering algorithms and provides network, dendrogram, and heat map views of the results. For example, protein-protein interaction data sets have been clustered to identify stable complexes, but scientists lack easily accessible tools to facilitate combined analyses of multiple data sets from different types of experiments. In the post-genomic era, the rapid increase in high-throughput data calls for computational tools capable of integrating data of diverse types and facilitating recognition of biologically meaningful patterns within them.
